| << Chapter < Page | Chapter >> Page > |
It should be noted that throughout the book, only scale-free graphs are allowed either:
We allow these exceptions so that our graphs will be in full compliance with the Cooper-Frieze model.
After generating a number of random graphs, we investigated a number of relationships between cell assemblies vs. other features in random graphs. We speculate on a few of these relationships here.
When we vary the probability that any given pair of vertices has an edge between them over a number of undirected, Bernoulli random graphs, we can see a number of distinct phases in the number of cell assemblies that these graphs will have, on average ( [link] ).

First, edge probability is insufficient to form even one assembly, then the number of assemblies increases to a peak as the graph becomes more dense. Past that peak, the closure of all tight sets converges to a single cell assembly. It would be interesting to find a method for predicting the probability at which this peak occurs, and then comparing that value to biology.
The Cooper-Frieze scale-free graph has no parameter to directly adjust edge density, but varying the related parameter, does have similar effects.
A fairly promising, although ostensibly not very accurate, indicator of the number of k-cores is the maximum eigenvalue of a graph's adjacency matrix ( [link] ).
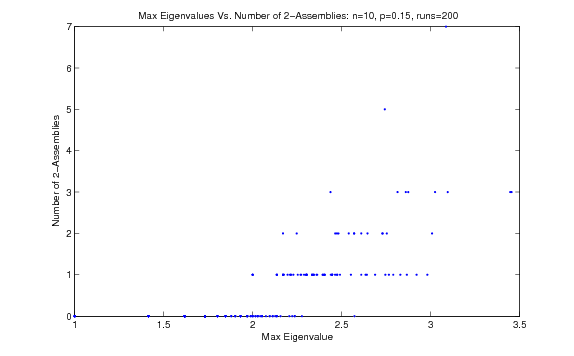
On undirected Bernoulli random graphs, the two variables appear to have a nontrivial, positive correlation. However, here we see that the method of graph construction is extremely important to any such observations, since on directed scale-free graphs, we see a relationship that is not nearly as clear ( [link] ).

Our approach to enumeration of cell assemblies in arbitrary graphs probably runs insufficiently fast to explore the problem as an end in itself. However, it does give us a useful tool to help us understand cell assemblies. We can now find an unlimited number of examples of cell assemblies which we may use as tools to explore general trends and gain insight into the structure of cell assemblies. Perhaps with enough work on the subject, we may find a viable way to understand the workings of the brain through random graph theory.

Notification Switch
Would you like to follow the 'The art of the pfug' conversation and receive update notifications?